Create a Graph Data Pipeline Using Python, Kafka and TigerGraph Kafka Loader
- Blog >
- Create a Graph Data Pipeline Using Python, Kafka and TigerGraph Kafka Loader
Originally posted on Medium by Kelley Brigman. Follow Kelley on Medium and Linkedin.
This article will discuss an efficient method for programmatically consuming datasets via REST API and loading them into TigerGraph using Kafka and TigerGraph Kafka Loader.
This article assumes that you have the following assets:
- A TigerGraph instance with access to the GSQL command line application, a secret and a token generated for that secret.
- Kafka and Zookeeper installed on a Linux server. If you need to install these, please refer to these instructions before continuing.
Overview
For this article I will create a Python application that will authenticate with a remote server, fetch a dataset via HTTP GET request, send the data records to a Kafka topic that is then consumed directly by a TigerGraph Kafka Load Job. The functions will be distributed across three severs: one for the application, one for Kafka and one for TigerGraph.
VPC Configuration
Because our Python application will use REST API methods to consume data from the UMLS, we need a NAT Gateway attached to a private subnet in our VPC. We also need a route from the subnet to the NAT Gateway.
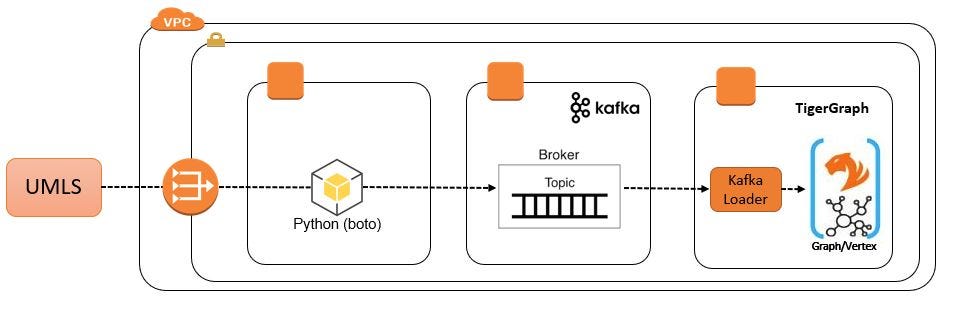
Simple VPC Configuration
VPC subnets gain access to a NAT Gateway via a route table. So a private subnet must be associated with a route table that has a path to the NAT Gateway. At minimum, the route table will look like this:
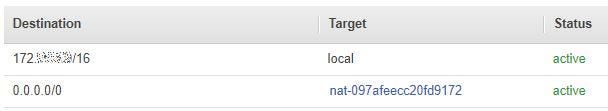
Route Table
Where 172.x.x.x/16 is the address for the VPC itself, 0.0.0.0/0 refers to the internet and nat-097afeecc20fd9172 is an AWS-generated name for the NAT Gateway. Together, these two route table entries grant traffic in the subnet permission to leave the subnet, move into local VPC space, then transit the NAT Gateway using the URL of the UMLS REST API. If you are unfamiliar with adding a NAT Gateway in your VPC, more information is available here.
Create Schema
Using TigerGraph’s Graph Studio, I have created a new vertex type named SNOMEDCT_Concept:
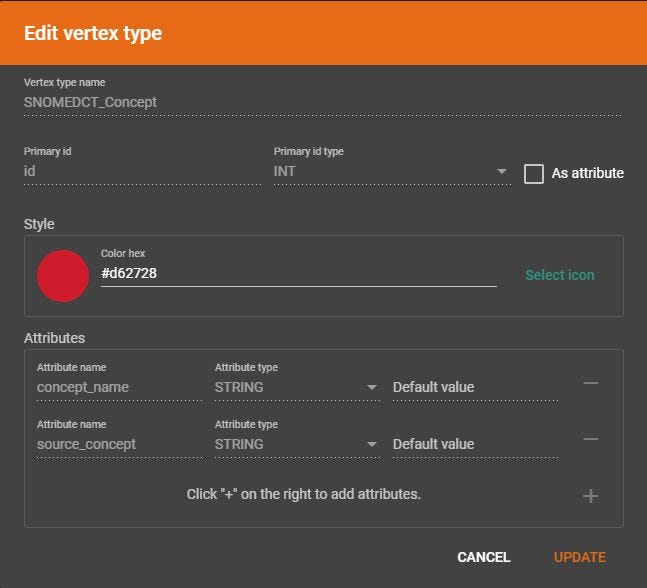
Create a Vertex Type
I have named the vertex SNOMEDCT_Concept. The primary id is id and the there are two string attributes, concept_name and source_concept. There is no need to upload a file or map data file to the vertex because we will be loading data via a Kafka topic.
Create a Kafka Topic
Log into the Kafka server as user kafka and create a new topic using the kafka-topics.sh shell script. I am using a single node for this demonstration, so replication factor is 1 and I will use 10 partitions.
~/kafka/bin/kafka-topics.sh --create --zookeeper localhost:2181 --replication-factor 1 --partitions 10 --topic SNOMEDCT_ConceptWe can see that the topic was created successfully using the same shell script.
~/kafka/bin/kafka-topics.sh --zookeeper localhost:2181 --listCreate a Loading Job
Log into the TigerGraph server as user tigergraph. Several steps are required before creating a TigerGraph Kafka Load Job. First, we need a data source configuration file. This is a small file that merely tells TigerGraph where to find the Kafka server. The value for the broker should be the same as the advertised.listeners value in the Kafka server configuration file (e.g. /home/kafka/kafka/config/server.properties). Don’t simply use “localhost”, the actual public IP address is preferred:
advertised.listeners=PLAINTEXT://161.xxx.xxx.xxx:9092
Multiple servers can be specified. Because my data pipeline application will be loading SNOMEDCT clinical concepts, I have named the file SNOMEDCT_Concept.json.
{
"broker": "161.xxx.xxx.xxx:9092"
}For convenience, I keep all the files related to Kafka loading jobs in the same location on the TigerGraph server, /home/tigergraph/datapipeline/load, but the file can reside anywhere that the TigerGraph user has read permission.
Next we create a partition configuration file. This file merely tells the TigerGraph Kafka Load Job which Kafka topic or topics to poll when loading data. I have named the file SNOMEDCT_Partition.json.
{
"topic": "SNOMEDCT_Concept"
}Next, we open the GSQL command line tool to create a Kafka data source that references the data source configuration file.
CREATE DATA_SOURCE KAFKA SNOMEDCT_Concept_Source = "/home/tigergraph/datapipeline/load/SNOMEDCT_Concept.json"
FOR GRAPH DataPipelineWe also grant permission for our data pipeline graph to use the loading job.
GRANT DATA_SOURCE SNOMEDCT_Concept_Source TO GRAPH DataPipelineNow we create a Kafka loading job.
CREATE LOADING JOB Load_SNOMEDCT_Concept
FOR GRAPH DataPipeline
{
DEFINE FILENAME f1 = "$SNOMEDCT_Concept_Source:SNOMEDCT_Partition.json";
LOAD f1 TO VERTEX SNOMEDCT_Concept VALUES ($0, $1, $2) USING SEPARATOR=”,”;
}And then run the loading job.
RUN LOADING JOB Load_SNOMEDCT_ConceptThe default action is to run the loading job indefinitely so that new records being added to the Kafka topic can be loaded immediately. Any record that already exists will be updated (based on the vertex id) and other records will be inserted. For each partition of this Kafka topic, we can see the number of loaded messages, the average speed, load duration and size of the data loaded so far. For example:
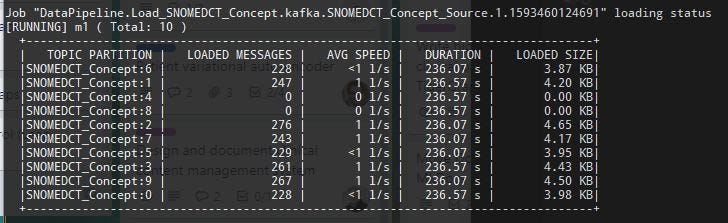
You could also run the loading job until it completes loading every record in the topic and then stop:
RUN LOADING JOB Load_SNOMEDCT_Concept EOF="true"More information about TigerGraph Kafka Loader is available here.
Consumer Data Via Rest API and Send to Kafka
Now all that remains is to write an application that gets data and sends the records to Kafka. I will be downloading SNOMEDCT Core Concepts from the Unified Medical Language Systems (UMLS). This service requires that members request an account and obtain an API key. Using this API key, we then generate a Ticket-granting Ticket (TGT) and a Service Ticket. I have encapsulated all the necessary logic for that process in a class called UmlsAuthentication. I will not discuss that logic in detail here but all the code is available on github here. Let’s take a look at the most important parts of the code.
First, we need a Kafka Producer object. This is our connection to the Kafka server and our means of publishing messages to a Kafka topic.
from kafka import KafkaProducer
kafka_producer = KafkaProducer(
bootstrap_servers=kafka_bootstrap_servers,
api_version=kafka_api_version
)kafka_bootstrap_servers is merely a list of IP addresses for the Kafka cluster. In this case, I am using a stand-alone server, so there is only one IP address. Make sure to specify port 9092 with the IP address (e.g.: 161.xxx.xxx.xxx:9092). I am using Kafka version 2.5.
Next, I get a list of configuration parameters from TigerGraph. Each record is contains everything I need to download the dataset, like the URL where the data can be obtained, the Kafka topic where it will be written, etc. The DataSetConfig class requires a TigerGraph secret and token. This is set in the initialization file for this project.
ds_config = DataSetConfig(data_set_name)Because there might be many, many records, we will download data from UMLS using paging. To start, I have set the page count variable to 100 records per page and the number of pages to 100. The effectively means that we will only get the first 1,000 records. So we start iterating:
for i in range(1, int(umls_api_fetch_page_count)):Next, we get a UMLS Service Token using my API key. A new Service Token is required for each HTTP GET, so we request one per iteration.
umls_auth = UmlsAuthentication(umls_api_key)Next, we create a URL that references the Service Ticket, page number and page size.
url = "{source_url}?ticket={service_ticket}&pageSize= {page_size}&pageNumber={page_no}".format(
source_url=ds_config.source_url,
service_ticket=umls_auth.service_ticket,
page_size=umls_api_fetch_page_size,
page_no=i
)Using this URL, we perform the HTTP request. If we get results, we put each record to the Kafka topic. Otherwise we show an error.
response = requests.get(url)
if response.status_code == 200:
data = json.loads(
response.content.decode(‘utf-8’)
).get(‘result’)
for d in data:
rec = “{0},{1},{2}”.format(
d.get(“ui”),
d.get(“name”),
d.get(“sourceConcept”)
)
kafka_producer.send(
ds_config.topic_name,
json.dumps(rec).encode(‘utf-8’)
)
else:
print(“UMLS did not return a valid reponse:
{0}”.format(response.status_code))Note that the record we send via the Kafka producer is a simple CSV string, however it should be UTF-8 encoded.
All that is left is to run our application and see the results streaming into TigerGraph. Using the Graph Explorer, I can see that I have many records. Each dot represents a vertex record.
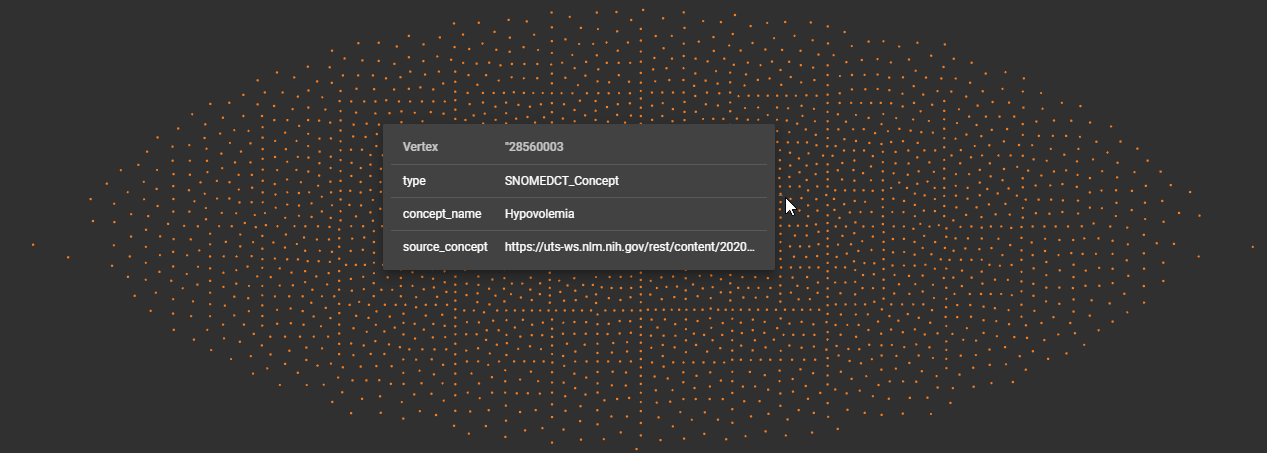
Graph Studio Explorer
And in the JSON results view, I can see that the concepts:
{
"v_id": ""428942009",
"v_type": "SNOMEDCT_Concept",
"attributes": {
"concept_name": "History of fall",
"source_concept": "https://uts-ws.nlm.nih.gov/rest/content/2020AA/source/SNOMEDCT_US/428942009/""}
},
{
"v_id": ""441935006",
"v_type": "SNOMEDCT_Concept",
"attributes": {
"concept_name": "Acquired absence of all teeth",
"source_concept": "https://uts-ws.nlm.nih.gov/rest/content/2020AA/source/SNOMEDCT_US/441935006/""}
}...
Conclusion
I hope you have found this article useful. While we covered only one simple example, this method can easily be expanded to include other, more complex cases like streaming sensor/IoT data, bank transactions or patient records.